fresh wraps fwapg and bcfishpass into
composable R functions for upstream/downstream network queries, habitat
model filtering, and waterbody traversal. All functions accept
table and cols parameters so the same
interface works across FWA base tables and bcfishpass model views.
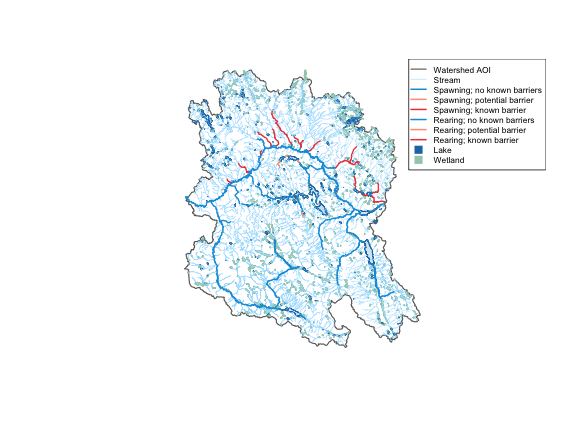
Here we work in the Neexdzii Kwa (Upper Bulkley River) watershed in the traditional territory of the Wet’suwet’en — querying upstream of the Neexdzii Kwa / Wedzin Kwa confluence to pull the full stream network, then zooming in to order 4+ coho rearing/spawning habitat from bcfishpass.
library(fresh)
library(sf)
#> Linking to GEOS 3.13.0, GDAL 3.8.5, PROJ 9.5.1; sf_use_s2() is TRUE
conn <- frs_db_conn()
blk <- 360873822
drm <- 166030.4
# Watershed AOI via frs_watershed_at_measure()
aoi <- frs_watershed_at_measure(conn, blk, drm)
# Full upstream network: streams, coho habitat, lakes, wetlands — one call
result <- frs_network(conn, blk, drm, tables = list(
streams = "whse_basemapping.fwa_stream_networks_sp",
co = list(
table = "bcfishpass.streams_co_vw",
cols = c("segmented_stream_id", "blue_line_key", "waterbody_key",
"gnis_name", "stream_order", "channel_width", "mapping_code",
"rearing", "spawning", "access", "geom"),
wscode_col = "wscode",
localcode_col = "localcode",
extra_where = "(s.rearing > 0 OR s.spawning > 0)"
),
lakes = "whse_basemapping.fwa_lakes_poly",
wetlands = "whse_basemapping.fwa_wetlands_poly"
))
streams_all <- result$streams
co_habitat <- result$co
lakes <- result$lakes
wetlands <- result$wetlands8876 stream segments in the full network narrow to 2133 coho habitat segments with rearing or spawning (orders 1, 2, 3, 4, 5, 6). 363 lakes and 1293 wetlands upstream.
reg <- gq::gq_reg_main()
cls <- gq::gq_tmap_classes(reg$layers$streams_salmon)
lake_style <- gq::gq_tmap_style(reg$layers$lake)
wetland_style <- gq::gq_tmap_style(reg$layers$wetland)
co_habitat$col <- cls$values[co_habitat$mapping_code]
co_habitat$col[is.na(co_habitat$col)] <- "#999999"
co_habitat$lwd <- ifelse(co_habitat$spawning > 0, 1.7, 1.0)
plot(st_geometry(aoi), col = NA, border = "grey40", lwd = 1.5, main = "")
if (nrow(wetlands) > 0) {
plot(st_geometry(wetlands), col = wetland_style$fill,
border = wetland_style$fill, add = TRUE)
}
if (nrow(lakes) > 0) {
plot(st_geometry(lakes), col = lake_style$fill,
border = lake_style$col, add = TRUE)
}
plot(st_geometry(streams_all), col = "#a9e0ff", lwd = 0.3, add = TRUE)
plot(st_geometry(co_habitat), col = co_habitat$col,
lwd = co_habitat$lwd, add = TRUE)
present <- names(cls$values) %in% unique(co_habitat$mapping_code)
legend(
"topright",
legend = c("Watershed AOI", "Stream", cls$labels[present], "Lake", "Wetland"),
col = c("grey40", "#a9e0ff", cls$values[present], lake_style$col, wetland_style$fill),
pch = c(NA, NA, rep(NA, sum(present)), 15, 15),
lwd = c(1.5, 0.8, rep(2, sum(present)), NA, NA),
pt.cex = c(NA, NA, rep(NA, sum(present)), 1.5, 1.5),
cex = 0.7, bg = "white"
)
See the function reference for the full API.
